Our team, which is led by Solon Pissis, works on algorithms and data structures for sequence analysis. We dive deep into the world of sequence analysis, crafting advanced algorithms and data structures to make sense of the vast amounts of sequential data all around us. Think of anything from the words you read every day to the staggering petabytes of genomic information that represent some of humanity's largest datasets.
Our team focuses on combinatorial pattern matching, an area where we design algorithms and data structures to manipulate strings. Our specific research topics include pattern matching, constructing indexes, comparing sequences, and finding regularities. While the developed techniques are vital for computational molecular biology—like analyzing DNA, RNA, or protein sequences—we also apply them to other areas that handle sequential data, such as data mining, data compression, and information retrieval.
Combinatorial pattern matching
Let us start by providing a brief overview of the combinatorial pattern matching topics.
In pattern matching, we are given a string x to pre-process in time that is proportional to the length of x, so that when we are given another longer string y, we can find all occurrences of x in y in time that is proportional to the length of y. In the following example we see two occurrences of string x in string y.
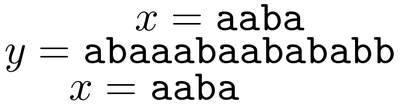
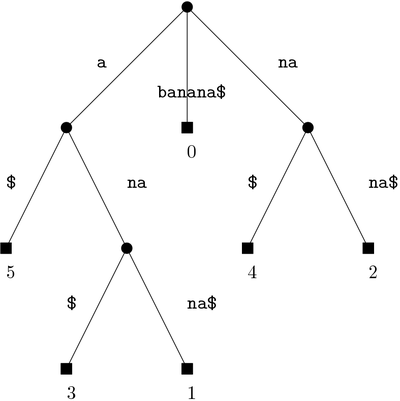
In indexing, we are given a string y to pre-process into a data structure in time that is proportional to the length of y, so that when we are given another shorter string x, we can find occurrences of x in y in time that is proportional to the length of x. In the following example we see an indexing data structure for string y = banana$. Every path from the root to a leaf node corresponds to a suffix of y. For instance, nana$ is the suffix starting at position 2 of y using a 0-base index.
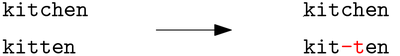
In sequence comparison, we are given two (or more) strings and the task is to compare them in order to infer their (dis)similarities. In the following example we see an alignment of two strings: kitchen and kitten.
In finding regularities, we are given a string x and the task is to find certain types of patterns repeating in x. In the following example we see that pattern aabbab repeats in string x = abaabbabbaaabbabbba.
Note the fundamental difference to pattern matching: here, the pattern is not given to us but we are rather asked to extract it from x.
Computational biology
Let us mention a few applications of combinatorial pattern matching in computational molecular biology:
- The first step of genome assembly via read mapping is to construct an indexing data structure over the reference genome. The first step of de novo genome assembly is to find common substrings between the reads to construct a de Bruijn or an overlap graph.
- The first step of inferring evolutionary relationships between sequences is the pairwise or multiple comparison of these sequences. The output of this step is used as the input to methods that reconstruct phylogenetic trees to depict these relationships.
- Inferring functional relationships between sequences is performed via comparing a sequence to a sequence database to detect regions of significant similarity or via extracting patterns (motifs), which are significantly overrepresented in a set of sequences.
Data mining
Combinatorial pattern matching methods are also applied in data mining, when the data to be analyzed are textual (sequential) and unstructured. For example, a string can represent the movement history of an individual, with each letter corresponding to a visited location; or an individual's purchasing history in a retailer, with each letter corresponding to a purchased product. Let us mention a few such applications:
- In pattern mining the task is to extract actionable patterns from large textual datasets; e.g., to extract the most frequent or the most significant patterns.
- In data sanitization the task is to disguise confidential patterns in large textual datasets while preserving the utility of these datasets.
- In document clustering the task is to compute term frequencies to reflect how important a term is to a document in a large collection of documents.
Researchers
Recent related research outputs
- Jialong Zhou, Ben Bals, Matei Tinca, Ai Guan, Panagiotis Charalampopoulos, Grigorios Loukides, Solon P. Pissis: Subtree Mode and Applications. ICDE 2026
- Ling Li, Daniel Gibney, Sharma V. Thankachan, Solon P. Pissis, Grigorios Loukides: Contextual Pattern Mining and Counting. ICDE 2026
- Pengxin Bian, Panagiotis Charalampopoulos, Lorraine Ayad, Manal Mohamed, Solon P. Pissis, Grigorios Loukides: Resilient Pattern Mining. ICDM 2025
- Lorraine A. K. Ayad, Grigorios Loukides, Solon P. Pissis: Text indexing for long patterns using locally consistent anchors. The VLDB Journal (2025)
- Ben Bals, Sebastiaan van Krieken, Solon Pissis, Leen Stougie, Hilde Verbeek. When is String Reconstruction using de Bruijn Graphs Hard? ESA 2025
Alumni
- Wiktor Zuba (Postdoc, Mar 2022 - Aug 2024)
- Esteban Gabory (Ph.D student, Oct 2021 - Aug 2024)
- Michelle Sweering (Ph.D student, Nov 2019 - Jan 2024)
- Giulia Bernardini (Postdoc, Jan 2021 - Dec 2021)
